Mol:FL5FAAGS0022
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 42 46 0 0 0 0 0 0 0 0999 V2000 | + | 42 46 0 0 0 0 0 0 0 0999 V2000 |
| − | -3.3489 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.3489 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.3489 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.3489 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7926 0.5054 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7926 0.5054 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2363 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2363 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2363 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2363 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7926 1.7902 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7926 1.7902 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6800 0.5054 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6800 0.5054 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1237 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1237 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1237 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1237 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6800 1.7902 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6800 1.7902 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6800 0.0046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6800 0.0046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5676 1.7900 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5676 1.7900 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0006 1.4627 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0006 1.4627 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5664 1.7900 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5664 1.7900 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5664 2.4447 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5664 2.4447 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0006 2.7721 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0006 2.7721 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5676 2.4447 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5676 2.4447 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7926 -0.1367 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7926 -0.1367 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.9050 1.7900 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.9050 1.7900 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1332 2.7720 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1332 2.7720 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5958 0.2988 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5958 0.2988 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1417 -0.7592 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 0.1417 -0.7592 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 0.7700 -1.1219 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.7700 -1.1219 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 0.5706 -0.4244 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 0.5706 -0.4244 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 0.7700 0.2773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7700 0.2773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1417 0.6401 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.1417 0.6401 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 0.3410 -0.0574 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.3410 -0.0574 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.1114 -0.0574 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1114 -0.0574 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3031 -1.3614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3031 -1.3614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6184 -1.6876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6184 -1.6876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1908 -0.7825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1908 -0.7825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2505 -1.6660 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 1.2505 -1.6660 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 1.7661 -2.3466 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 1.7661 -2.3466 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.5086 -2.0579 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.5086 -2.0579 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.2251 -2.0502 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.2251 -2.0502 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.7045 -1.5294 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 2.7045 -1.5294 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.0549 -1.8723 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0549 -1.8723 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.9623 -2.3858 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.9623 -2.3858 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.9341 -2.7721 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9341 -2.7721 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2163 -2.1824 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2163 -2.1824 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.4614 -0.8759 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.4614 -0.8759 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2739 0.1064 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2739 0.1064 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 1 19 1 0 0 0 0 | + | 1 19 1 0 0 0 0 |
| − | 15 20 1 0 0 0 0 | + | 15 20 1 0 0 0 0 |
| − | 21 8 1 0 0 0 0 | + | 21 8 1 0 0 0 0 |
| − | 23 22 1 1 0 0 0 | + | 23 22 1 1 0 0 0 |
| − | 23 24 1 0 0 0 0 | + | 23 24 1 0 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 1 1 0 0 0 | + | 26 27 1 1 0 0 0 |
| − | 27 22 1 1 0 0 0 | + | 27 22 1 1 0 0 0 |
| − | 27 28 1 0 0 0 0 | + | 27 28 1 0 0 0 0 |
| − | 22 29 1 0 0 0 0 | + | 22 29 1 0 0 0 0 |
| − | 23 30 1 0 0 0 0 | + | 23 30 1 0 0 0 0 |
| − | 24 31 1 0 0 0 0 | + | 24 31 1 0 0 0 0 |
| − | 21 26 1 0 0 0 0 | + | 21 26 1 0 0 0 0 |
| − | 32 33 1 1 0 0 0 | + | 32 33 1 1 0 0 0 |
| − | 33 34 1 1 0 0 0 | + | 33 34 1 1 0 0 0 |
| − | 35 34 1 1 0 0 0 | + | 35 34 1 1 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 37 1 0 0 0 0 | + | 36 37 1 0 0 0 0 |
| − | 37 32 1 0 0 0 0 | + | 37 32 1 0 0 0 0 |
| − | 33 38 1 0 0 0 0 | + | 33 38 1 0 0 0 0 |
| − | 34 39 1 0 0 0 0 | + | 34 39 1 0 0 0 0 |
| − | 35 40 1 0 0 0 0 | + | 35 40 1 0 0 0 0 |
| − | 32 30 1 0 0 0 0 | + | 32 30 1 0 0 0 0 |
| − | 36 41 1 0 0 0 0 | + | 36 41 1 0 0 0 0 |
| − | 41 42 1 0 0 0 0 | + | 41 42 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 41 42 | + | M SAL 1 2 41 42 |
| − | M SBL 1 1 45 | + | M SBL 1 1 45 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SVB 1 45 3.0028 -1.4895 | + | M SVB 1 45 3.0028 -1.4895 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FAAGS0022 | + | ID FL5FAAGS0022 |
| − | KNApSAcK_ID C00005172 | + | KNApSAcK_ID C00005172 |
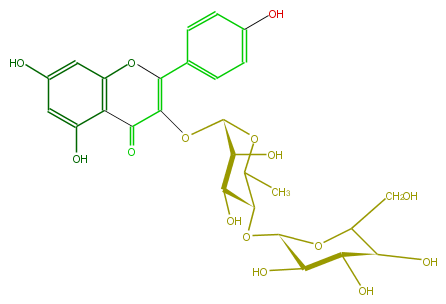
| − | NAME Multiflorin B | + | NAME Multiflorin B |
| − | CAS_RN 52657-01-9 | + | CAS_RN 52657-01-9 |
| − | FORMULA C27H30O15 | + | FORMULA C27H30O15 |
| − | EXACTMASS 594.15847029 | + | EXACTMASS 594.15847029 |
| − | AVERAGEMASS 594.5181 | + | AVERAGEMASS 594.5181 |
| − | SMILES c(O)(c5)cc(c(c35)C(=O)C(=C(c(c4)ccc(c4)O)O3)O[C@@H](O1)[C@@H](O)[C@H](O)[C@H](O[C@@H](O2)[C@H](O)[C@H](O)[C@@H](C2CO)O)C(C)1)O | + | SMILES c(O)(c5)cc(c(c35)C(=O)C(=C(c(c4)ccc(c4)O)O3)O[C@@H](O1)[C@@H](O)[C@H](O)[C@H](O[C@@H](O2)[C@H](O)[C@H](O)[C@@H](C2CO)O)C(C)1)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
42 46 0 0 0 0 0 0 0 0999 V2000
-3.3489 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.3489 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7926 0.5054 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2363 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2363 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7926 1.7902 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6800 0.5054 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1237 0.8266 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1237 1.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6800 1.7902 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.6800 0.0046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.5676 1.7900 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0006 1.4627 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5664 1.7900 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5664 2.4447 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0006 2.7721 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5676 2.4447 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7926 -0.1367 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.9050 1.7900 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1332 2.7720 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.5958 0.2988 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.1417 -0.7592 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
0.7700 -1.1219 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
0.5706 -0.4244 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.7700 0.2773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.1417 0.6401 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
0.3410 -0.0574 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.1114 -0.0574 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.3031 -1.3614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6184 -1.6876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1908 -0.7825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2505 -1.6660 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
1.7661 -2.3466 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.5086 -2.0579 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.2251 -2.0502 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.7045 -1.5294 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.0549 -1.8723 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.9623 -2.3858 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.9341 -2.7721 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2163 -2.1824 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.4614 -0.8759 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2739 0.1064 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
1 19 1 0 0 0 0
15 20 1 0 0 0 0
21 8 1 0 0 0 0
23 22 1 1 0 0 0
23 24 1 0 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 27 1 1 0 0 0
27 22 1 1 0 0 0
27 28 1 0 0 0 0
22 29 1 0 0 0 0
23 30 1 0 0 0 0
24 31 1 0 0 0 0
21 26 1 0 0 0 0
32 33 1 1 0 0 0
33 34 1 1 0 0 0
35 34 1 1 0 0 0
35 36 1 0 0 0 0
36 37 1 0 0 0 0
37 32 1 0 0 0 0
33 38 1 0 0 0 0
34 39 1 0 0 0 0
35 40 1 0 0 0 0
32 30 1 0 0 0 0
36 41 1 0 0 0 0
41 42 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 41 42
M SBL 1 1 45
M SMT 1 CH2OH
M SVB 1 45 3.0028 -1.4895
S SKP 8
ID FL5FAAGS0022
KNApSAcK_ID C00005172
NAME Multiflorin B
CAS_RN 52657-01-9
FORMULA C27H30O15
EXACTMASS 594.15847029
AVERAGEMASS 594.5181
SMILES c(O)(c5)cc(c(c35)C(=O)C(=C(c(c4)ccc(c4)O)O3)O[C@@H](O1)[C@@H](O)[C@H](O)[C@H](O[C@@H](O2)[C@H](O)[C@H](O)[C@@H](C2CO)O)C(C)1)O
M END