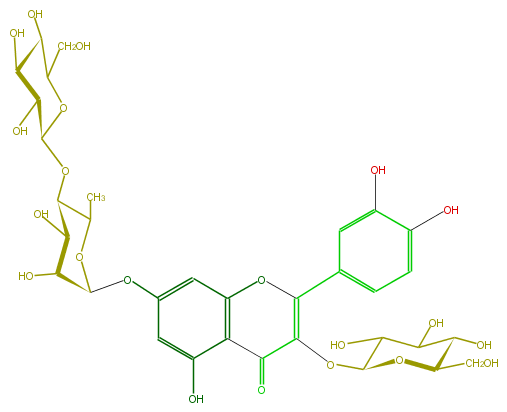
Mol:FL5FACGL0049
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 54 59 0 0 0 0 0 0 0 0999 V2000 | + | 54 59 0 0 0 0 0 0 0 0999 V2000 |
| − | -2.1853 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1853 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1853 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1853 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4706 -3.1676 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4706 -3.1676 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7560 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7560 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7560 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7560 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4706 -1.5172 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4706 -1.5172 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0414 -3.1676 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0414 -3.1676 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6733 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6733 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6733 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6733 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0414 -1.5172 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0414 -1.5172 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0414 -3.8111 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0414 -3.8111 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5732 -1.3584 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5732 -1.3584 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3016 -1.7789 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3016 -1.7789 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0300 -1.3584 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0300 -1.3584 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0300 -0.5173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0300 -0.5173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3016 -0.0969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3016 -0.0969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5732 -0.5173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5732 -0.5173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4706 -3.9926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4706 -3.9926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8095 -0.0673 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8095 -0.0673 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8211 -1.5172 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8211 -1.5172 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3969 -3.2811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3969 -3.2811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3016 0.7439 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3016 0.7439 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2807 -1.4107 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2807 -1.4107 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5995 -1.8039 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5995 -1.8039 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.8158 -1.0477 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8158 -1.0477 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5995 -0.2868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5995 -0.2868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2807 0.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2807 0.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0646 -0.6498 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0646 -0.6498 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.1119 0.7367 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.1119 0.7367 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5995 0.2040 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5995 0.2040 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.6760 -0.1570 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.6760 -0.1570 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8547 -1.4425 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8547 -1.4425 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.6145 3.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.6145 3.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.1184 2.7912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.1184 2.7912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.6680 2.1995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.6680 2.1995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.6002 1.3928 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.6002 1.3928 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.1403 2.0400 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.1403 2.0400 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.4921 2.6571 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.4921 2.6571 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8001 3.9926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8001 3.9926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.1803 3.5917 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.1803 3.5917 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4559 -2.7330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4559 -2.7330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0693 -3.4025 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0693 -3.4025 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8125 -3.1900 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8125 -3.1900 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5602 -3.4025 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5602 -3.4025 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.9467 -2.7330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.9467 -2.7330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2036 -2.9454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2036 -2.9454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6158 -2.8650 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6158 -2.8650 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5094 -2.4155 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5094 -2.4155 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.5056 -2.8709 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.5056 -2.8709 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.1250 1.5691 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.1250 1.5691 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.1864 3.3268 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.1864 3.3268 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.2687 2.7969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2687 2.7969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2625 -3.2458 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2625 -3.2458 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.1803 -3.7757 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.1803 -3.7757 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 19 15 1 0 0 0 0 | + | 19 15 1 0 0 0 0 |
| − | 20 1 1 0 0 0 0 | + | 20 1 1 0 0 0 0 |
| − | 21 8 1 0 0 0 0 | + | 21 8 1 0 0 0 0 |
| − | 4 3 1 0 0 0 0 | + | 4 3 1 0 0 0 0 |
| − | 16 22 1 0 0 0 0 | + | 16 22 1 0 0 0 0 |
| − | 24 23 1 1 0 0 0 | + | 24 23 1 1 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 28 1 1 0 0 0 | + | 27 28 1 1 0 0 0 |
| − | 28 23 1 1 0 0 0 | + | 28 23 1 1 0 0 0 |
| − | 27 29 1 0 0 0 0 | + | 27 29 1 0 0 0 0 |
| − | 26 30 1 0 0 0 0 | + | 26 30 1 0 0 0 0 |
| − | 28 31 1 0 0 0 0 | + | 28 31 1 0 0 0 0 |
| − | 23 32 1 0 0 0 0 | + | 23 32 1 0 0 0 0 |
| − | 20 24 1 0 0 0 0 | + | 20 24 1 0 0 0 0 |
| − | 33 34 1 1 0 0 0 | + | 33 34 1 1 0 0 0 |
| − | 34 35 1 1 0 0 0 | + | 34 35 1 1 0 0 0 |
| − | 36 35 1 1 0 0 0 | + | 36 35 1 1 0 0 0 |
| − | 36 37 1 0 0 0 0 | + | 36 37 1 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | 38 33 1 0 0 0 0 | + | 38 33 1 0 0 0 0 |
| − | 33 39 1 0 0 0 0 | + | 33 39 1 0 0 0 0 |
| − | 34 40 1 0 0 0 0 | + | 34 40 1 0 0 0 0 |
| − | 36 29 1 0 0 0 0 | + | 36 29 1 0 0 0 0 |
| − | 41 42 1 0 0 0 0 | + | 41 42 1 0 0 0 0 |
| − | 42 43 1 1 0 0 0 | + | 42 43 1 1 0 0 0 |
| − | 43 44 1 1 0 0 0 | + | 43 44 1 1 0 0 0 |
| − | 45 44 1 1 0 0 0 | + | 45 44 1 1 0 0 0 |
| − | 45 46 1 0 0 0 0 | + | 45 46 1 0 0 0 0 |
| − | 46 41 1 0 0 0 0 | + | 46 41 1 0 0 0 0 |
| − | 41 47 1 0 0 0 0 | + | 41 47 1 0 0 0 0 |
| − | 46 48 1 0 0 0 0 | + | 46 48 1 0 0 0 0 |
| − | 45 49 1 0 0 0 0 | + | 45 49 1 0 0 0 0 |
| − | 42 21 1 0 0 0 0 | + | 42 21 1 0 0 0 0 |
| − | 35 50 1 0 0 0 0 | + | 35 50 1 0 0 0 0 |
| − | 51 52 1 0 0 0 0 | + | 51 52 1 0 0 0 0 |
| − | 38 51 1 0 0 0 0 | + | 38 51 1 0 0 0 0 |
| − | 53 54 1 0 0 0 0 | + | 53 54 1 0 0 0 0 |
| − | 44 53 1 0 0 0 0 | + | 44 53 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 51 52 | + | M SAL 1 2 51 52 |
| − | M SBL 1 1 57 | + | M SBL 1 1 57 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SBV 1 57 -0.3057 -0.6697 | + | M SBV 1 57 -0.3057 -0.6697 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 53 54 | + | M SAL 2 2 53 54 |
| − | M SBL 2 1 59 | + | M SBL 2 1 59 |
| − | M SMT 2 CH2OH | + | M SMT 2 CH2OH |
| − | M SBV 2 59 -0.7024 -0.1567 | + | M SBV 2 59 -0.7024 -0.1567 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL5FACGL0049 | + | ID FL5FACGL0049 |
| − | FORMULA C33H40O21 | + | FORMULA C33H40O21 |
| − | EXACTMASS 772.206208342 | + | EXACTMASS 772.206208342 |
| − | AVERAGEMASS 772.6581 | + | AVERAGEMASS 772.6581 |
| − | SMILES C(=C5c(c6)cc(c(c6)O)O)(C(=O)c(c4O5)c(cc(c4)OC(C(O)2)OC(C)C(OC(C(O)3)OC(CO)C(O)C3O)C2O)O)OC(O1)C(O)C(C(O)C1CO)O | + | SMILES C(=C5c(c6)cc(c(c6)O)O)(C(=O)c(c4O5)c(cc(c4)OC(C(O)2)OC(C)C(OC(C(O)3)OC(CO)C(O)C3O)C2O)O)OC(O1)C(O)C(C(O)C1CO)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
54 59 0 0 0 0 0 0 0 0999 V2000
-2.1853 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1853 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4706 -3.1676 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7560 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7560 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4706 -1.5172 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0414 -3.1676 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6733 -2.7551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6733 -1.9298 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0414 -1.5172 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.0414 -3.8111 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5732 -1.3584 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.3016 -1.7789 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0300 -1.3584 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0300 -0.5173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.3016 -0.0969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5732 -0.5173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4706 -3.9926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8095 -0.0673 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.8211 -1.5172 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.3969 -3.2811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.3016 0.7439 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2807 -1.4107 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.5995 -1.8039 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.8158 -1.0477 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.5995 -0.2868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.2807 0.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.0646 -0.6498 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.1119 0.7367 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.5995 0.2040 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.6760 -0.1570 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.8547 -1.4425 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.6145 3.4690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.1184 2.7912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.6680 2.1995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.6002 1.3928 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.1403 2.0400 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.4921 2.6571 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8001 3.9926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.1803 3.5917 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.4559 -2.7330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0693 -3.4025 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.8125 -3.1900 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.5602 -3.4025 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.9467 -2.7330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2036 -2.9454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.6158 -2.8650 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.5094 -2.4155 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.5056 -2.8709 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.1250 1.5691 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.1864 3.3268 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.2687 2.7969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2625 -3.2458 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.1803 -3.7757 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
19 15 1 0 0 0 0
20 1 1 0 0 0 0
21 8 1 0 0 0 0
4 3 1 0 0 0 0
16 22 1 0 0 0 0
24 23 1 1 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 27 1 0 0 0 0
27 28 1 1 0 0 0
28 23 1 1 0 0 0
27 29 1 0 0 0 0
26 30 1 0 0 0 0
28 31 1 0 0 0 0
23 32 1 0 0 0 0
20 24 1 0 0 0 0
33 34 1 1 0 0 0
34 35 1 1 0 0 0
36 35 1 1 0 0 0
36 37 1 0 0 0 0
37 38 1 0 0 0 0
38 33 1 0 0 0 0
33 39 1 0 0 0 0
34 40 1 0 0 0 0
36 29 1 0 0 0 0
41 42 1 0 0 0 0
42 43 1 1 0 0 0
43 44 1 1 0 0 0
45 44 1 1 0 0 0
45 46 1 0 0 0 0
46 41 1 0 0 0 0
41 47 1 0 0 0 0
46 48 1 0 0 0 0
45 49 1 0 0 0 0
42 21 1 0 0 0 0
35 50 1 0 0 0 0
51 52 1 0 0 0 0
38 51 1 0 0 0 0
53 54 1 0 0 0 0
44 53 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 51 52
M SBL 1 1 57
M SMT 1 CH2OH
M SBV 1 57 -0.3057 -0.6697
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 53 54
M SBL 2 1 59
M SMT 2 CH2OH
M SBV 2 59 -0.7024 -0.1567
S SKP 5
ID FL5FACGL0049
FORMULA C33H40O21
EXACTMASS 772.206208342
AVERAGEMASS 772.6581
SMILES C(=C5c(c6)cc(c(c6)O)O)(C(=O)c(c4O5)c(cc(c4)OC(C(O)2)OC(C)C(OC(C(O)3)OC(CO)C(O)C3O)C2O)O)OC(O1)C(O)C(C(O)C1CO)O
M END