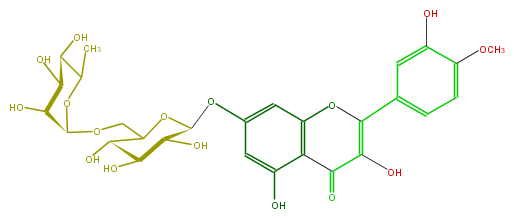
Mol:FL5FAEGS0004
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 44 48 0 0 0 0 0 0 0 0999 V2000 | + | 44 48 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.6030 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6030 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6030 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6030 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1115 -1.5430 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1115 -1.5430 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8259 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8259 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8259 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8259 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1115 0.1070 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1115 0.1070 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5404 -1.5430 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5404 -1.5430 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2549 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2549 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2549 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2549 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5404 0.1070 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5404 0.1070 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5404 -2.1861 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5404 -2.1861 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.1545 0.2657 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.1545 0.2657 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8826 -0.1546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8826 -0.1546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.6108 0.2657 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.6108 0.2657 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.6108 1.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.6108 1.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8826 1.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8826 1.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.1545 1.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.1545 1.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1115 -2.3676 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1115 -2.3676 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4564 0.1873 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4564 0.1873 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8826 2.3676 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8826 2.3676 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0034 -1.5626 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0034 -1.5626 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.9622 -0.8153 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.9622 -0.8153 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.2308 -1.2877 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2308 -1.2877 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6200 -0.7520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.6200 -0.7520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.9362 -0.4948 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.9362 -0.4948 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6166 -0.1773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.6166 -0.1773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.1196 -0.7324 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.1196 -0.7324 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6414 -0.3710 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6414 -0.3710 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.4765 -1.2455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.4765 -1.2455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.8573 -1.4481 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8573 -1.4481 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8176 -0.8976 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8176 -0.8976 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.5454 -0.0768 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.5454 -0.0768 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.0224 -0.5998 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.0224 -0.5998 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.0345 0.1399 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.0345 0.1399 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.6531 0.7784 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.6531 0.7784 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.1760 1.3014 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.1760 1.3014 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.1638 0.5617 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.1638 0.5617 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2810 -0.5543 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2810 -0.5543 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.7858 1.7720 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.7858 1.7720 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.5416 1.5004 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.5416 1.5004 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.6824 1.2311 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.6824 1.2311 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -6.1854 0.0537 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -6.1854 0.0537 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.2677 1.4933 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.2677 1.4933 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 6.1854 0.9636 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 6.1854 0.9636 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 4 3 1 0 0 0 0 | + | 4 3 1 0 0 0 0 |
| − | 1 19 1 0 0 0 0 | + | 1 19 1 0 0 0 0 |
| − | 16 20 1 0 0 0 0 | + | 16 20 1 0 0 0 0 |
| − | 21 8 1 0 0 0 0 | + | 21 8 1 0 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 25 24 1 1 0 0 0 | + | 25 24 1 1 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 22 1 0 0 0 0 | + | 27 22 1 0 0 0 0 |
| − | 27 28 1 0 0 0 0 | + | 27 28 1 0 0 0 0 |
| − | 22 29 1 0 0 0 0 | + | 22 29 1 0 0 0 0 |
| − | 23 30 1 0 0 0 0 | + | 23 30 1 0 0 0 0 |
| − | 24 31 1 0 0 0 0 | + | 24 31 1 0 0 0 0 |
| − | 33 32 1 1 0 0 0 | + | 33 32 1 1 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 37 1 1 0 0 0 | + | 36 37 1 1 0 0 0 |
| − | 37 32 1 1 0 0 0 | + | 37 32 1 1 0 0 0 |
| − | 33 38 1 0 0 0 0 | + | 33 38 1 0 0 0 0 |
| − | 38 28 1 0 0 0 0 | + | 38 28 1 0 0 0 0 |
| − | 36 39 1 0 0 0 0 | + | 36 39 1 0 0 0 0 |
| − | 35 40 1 0 0 0 0 | + | 35 40 1 0 0 0 0 |
| − | 37 41 1 0 0 0 0 | + | 37 41 1 0 0 0 0 |
| − | 32 42 1 0 0 0 0 | + | 32 42 1 0 0 0 0 |
| − | 25 19 1 0 0 0 0 | + | 25 19 1 0 0 0 0 |
| − | 43 44 1 0 0 0 0 | + | 43 44 1 0 0 0 0 |
| − | 15 43 1 0 0 0 0 | + | 15 43 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 43 44 | + | M SAL 1 2 43 44 |
| − | M SBL 1 1 48 | + | M SBL 1 1 48 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SBV 1 48 -0.6569 -0.3868 | + | M SBV 1 48 -0.6569 -0.3868 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL5FAEGS0004 | + | ID FL5FAEGS0004 |
| − | FORMULA C28H32O16 | + | FORMULA C28H32O16 |
| − | EXACTMASS 624.1690349759999 | + | EXACTMASS 624.1690349759999 |
| − | AVERAGEMASS 624.54408 | + | AVERAGEMASS 624.54408 |
| − | SMILES COc(c5)c(O)cc(c5)C(=C(O)4)Oc(c3C4=O)cc(cc3O)OC(O1)C(O)C(O)C(O)C(COC(O2)C(C(C(O)C(C)2)O)O)1 | + | SMILES COc(c5)c(O)cc(c5)C(=C(O)4)Oc(c3C4=O)cc(cc3O)OC(O1)C(O)C(O)C(O)C(COC(O2)C(C(C(O)C(C)2)O)O)1 |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
44 48 0 0 0 0 0 0 0 0999 V2000
-0.6030 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6030 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1115 -1.5430 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8259 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8259 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1115 0.1070 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5404 -1.5430 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2549 -1.1304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2549 -0.3055 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5404 0.1070 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5404 -2.1861 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.1545 0.2657 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.8826 -0.1546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.6108 0.2657 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.6108 1.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.8826 1.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.1545 1.1066 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1115 -2.3676 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.4564 0.1873 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8826 2.3676 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.0034 -1.5626 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.9622 -0.8153 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.2308 -1.2877 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.6200 -0.7520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.9362 -0.4948 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.6166 -0.1773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.1196 -0.7324 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.6414 -0.3710 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.4765 -1.2455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.8573 -1.4481 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.8176 -0.8976 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.5454 -0.0768 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.0224 -0.5998 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.0345 0.1399 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.6531 0.7784 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.1760 1.3014 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.1638 0.5617 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.2810 -0.5543 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.7858 1.7720 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.5416 1.5004 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.6824 1.2311 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-6.1854 0.0537 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.2677 1.4933 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
6.1854 0.9636 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
4 3 1 0 0 0 0
1 19 1 0 0 0 0
16 20 1 0 0 0 0
21 8 1 0 0 0 0
22 23 1 1 0 0 0
23 24 1 1 0 0 0
25 24 1 1 0 0 0
25 26 1 0 0 0 0
26 27 1 0 0 0 0
27 22 1 0 0 0 0
27 28 1 0 0 0 0
22 29 1 0 0 0 0
23 30 1 0 0 0 0
24 31 1 0 0 0 0
33 32 1 1 0 0 0
33 34 1 0 0 0 0
34 35 1 0 0 0 0
35 36 1 0 0 0 0
36 37 1 1 0 0 0
37 32 1 1 0 0 0
33 38 1 0 0 0 0
38 28 1 0 0 0 0
36 39 1 0 0 0 0
35 40 1 0 0 0 0
37 41 1 0 0 0 0
32 42 1 0 0 0 0
25 19 1 0 0 0 0
43 44 1 0 0 0 0
15 43 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 43 44
M SBL 1 1 48
M SMT 1 OCH3
M SBV 1 48 -0.6569 -0.3868
S SKP 5
ID FL5FAEGS0004
FORMULA C28H32O16
EXACTMASS 624.1690349759999
AVERAGEMASS 624.54408
SMILES COc(c5)c(O)cc(c5)C(=C(O)4)Oc(c3C4=O)cc(cc3O)OC(O1)C(O)C(O)C(O)C(COC(O2)C(C(C(O)C(C)2)O)O)1
M END