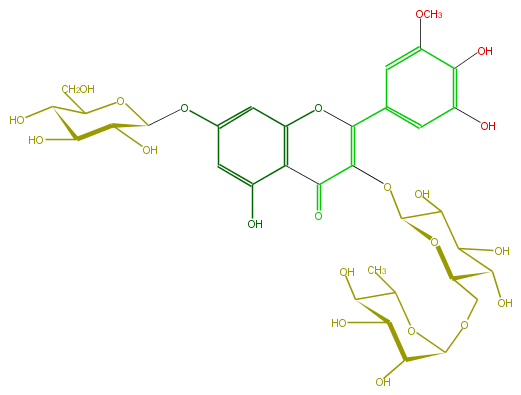
Mol:FL5FAHGL0004
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 56 61 0 0 0 0 0 0 0 0999 V2000 | + | 56 61 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.7143 0.9256 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7143 0.9256 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7143 0.1006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7143 0.1006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0002 -0.3118 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0002 -0.3118 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7146 0.1005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7146 0.1005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7144 0.9255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7144 0.9255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0000 1.3380 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0000 1.3380 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4291 -0.3118 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4291 -0.3118 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1434 0.1005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1434 0.1005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1433 0.9255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1433 0.9255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4288 1.3381 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4288 1.3381 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4291 -0.9550 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4291 -0.9550 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8576 1.3379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8576 1.3379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5857 0.9175 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5857 0.9175 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3138 1.3379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3138 1.3379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3138 2.1786 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3138 2.1786 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5858 2.5991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5858 2.5991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8576 2.1786 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8576 2.1786 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4285 1.3379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4285 1.3379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8985 -0.3385 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8985 -0.3385 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0003 -1.1365 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0003 -1.1365 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0553 -2.6093 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0553 -2.6093 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2062 -2.7590 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2062 -2.7590 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.9206 -3.2419 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9206 -3.2419 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2753 -4.0332 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2753 -4.0332 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.1245 -3.8836 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.1245 -3.8836 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.4100 -3.4005 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.4100 -3.4005 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5465 -2.0999 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5465 -2.0999 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0418 -0.8823 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0418 -0.8823 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.1926 -1.0319 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.1926 -1.0319 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.9071 -1.5148 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.9071 -1.5148 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2618 -2.3061 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2618 -2.3061 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.1110 -2.1565 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.1110 -2.1565 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3965 -1.6735 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3965 -1.6735 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5525 -0.4933 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5525 -0.4933 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.2560 -1.7008 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.2560 -1.7008 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.3240 -2.8092 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.3240 -2.8092 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.7919 -2.7480 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.7919 -2.7480 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.5375 -3.2782 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.5375 -3.2782 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9587 -2.1419 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9587 -2.1419 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9354 -3.2364 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9354 -3.2364 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7180 -4.5008 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7180 -4.5008 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.8930 2.5663 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.8930 2.5663 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.9672 0.9665 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.9672 0.9665 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2508 1.3499 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2508 1.3499 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.7300 0.6625 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.7300 0.6625 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.9803 0.9540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.9803 0.9540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1988 0.9454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1988 0.9454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7825 1.4877 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7825 1.4877 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5484 1.2127 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5484 1.2127 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.9190 1.0770 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.9190 1.0770 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.5088 0.6534 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.5088 0.6534 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2397 0.4379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2397 0.4379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5858 3.3485 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5858 3.3485 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.1527 4.5008 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.1527 4.5008 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0709 1.7353 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0709 1.7353 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.3240 2.0167 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.3240 2.0167 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 1 18 1 0 0 0 0 | + | 1 18 1 0 0 0 0 |
| − | 8 19 1 0 0 0 0 | + | 8 19 1 0 0 0 0 |
| − | 3 20 1 0 0 0 0 | + | 3 20 1 0 0 0 0 |
| − | 21 22 1 0 0 0 0 | + | 21 22 1 0 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 25 24 1 1 0 0 0 | + | 25 24 1 1 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 21 1 0 0 0 0 | + | 26 21 1 0 0 0 0 |
| − | 21 27 1 0 0 0 0 | + | 21 27 1 0 0 0 0 |
| − | 28 29 1 0 0 0 0 | + | 28 29 1 0 0 0 0 |
| − | 29 30 1 1 0 0 0 | + | 29 30 1 1 0 0 0 |
| − | 30 31 1 1 0 0 0 | + | 30 31 1 1 0 0 0 |
| − | 32 31 1 1 0 0 0 | + | 32 31 1 1 0 0 0 |
| − | 32 33 1 0 0 0 0 | + | 32 33 1 0 0 0 0 |
| − | 33 28 1 0 0 0 0 | + | 33 28 1 0 0 0 0 |
| − | 28 34 1 0 0 0 0 | + | 28 34 1 0 0 0 0 |
| − | 33 35 1 0 0 0 0 | + | 33 35 1 0 0 0 0 |
| − | 32 36 1 0 0 0 0 | + | 32 36 1 0 0 0 0 |
| − | 31 37 1 0 0 0 0 | + | 31 37 1 0 0 0 0 |
| − | 25 38 1 0 0 0 0 | + | 25 38 1 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | 22 39 1 0 0 0 0 | + | 22 39 1 0 0 0 0 |
| − | 23 40 1 0 0 0 0 | + | 23 40 1 0 0 0 0 |
| − | 24 41 1 0 0 0 0 | + | 24 41 1 0 0 0 0 |
| − | 29 19 1 0 0 0 0 | + | 29 19 1 0 0 0 0 |
| − | 42 15 1 0 0 0 0 | + | 42 15 1 0 0 0 0 |
| − | 14 43 1 0 0 0 0 | + | 14 43 1 0 0 0 0 |
| − | 44 45 1 1 0 0 0 | + | 44 45 1 1 0 0 0 |
| − | 45 46 1 1 0 0 0 | + | 45 46 1 1 0 0 0 |
| − | 47 46 1 1 0 0 0 | + | 47 46 1 1 0 0 0 |
| − | 47 48 1 0 0 0 0 | + | 47 48 1 0 0 0 0 |
| − | 48 49 1 0 0 0 0 | + | 48 49 1 0 0 0 0 |
| − | 49 44 1 0 0 0 0 | + | 49 44 1 0 0 0 0 |
| − | 44 50 1 0 0 0 0 | + | 44 50 1 0 0 0 0 |
| − | 45 51 1 0 0 0 0 | + | 45 51 1 0 0 0 0 |
| − | 46 52 1 0 0 0 0 | + | 46 52 1 0 0 0 0 |
| − | 47 18 1 0 0 0 0 | + | 47 18 1 0 0 0 0 |
| − | 53 54 1 0 0 0 0 | + | 53 54 1 0 0 0 0 |
| − | 16 53 1 0 0 0 0 | + | 16 53 1 0 0 0 0 |
| − | 55 56 1 0 0 0 0 | + | 55 56 1 0 0 0 0 |
| − | 49 55 1 0 0 0 0 | + | 49 55 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 53 54 | + | M SAL 1 2 53 54 |
| − | M SBL 1 1 59 | + | M SBL 1 1 59 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SBV 1 59 0.0000 -0.7494 | + | M SBV 1 59 0.0000 -0.7494 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 55 56 | + | M SAL 2 2 55 56 |
| − | M SBL 2 1 61 | + | M SBL 2 1 61 |
| − | M SMT 2 ^ CH2OH | + | M SMT 2 ^ CH2OH |
| − | M SBV 2 61 0.5225 -0.5225 | + | M SBV 2 61 0.5225 -0.5225 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL5FAHGL0004 | + | ID FL5FAHGL0004 |
| − | FORMULA C34H42O22 | + | FORMULA C34H42O22 |
| − | EXACTMASS 802.216773028 | + | EXACTMASS 802.216773028 |
| − | AVERAGEMASS 802.68408 | + | AVERAGEMASS 802.68408 |
| − | SMILES C(C(O)6)(OC(C(O)C(O)6)Oc(c1)cc(c(C3=O)c1OC(=C(OC(C5O)OC(C(C(O)5)O)COC(C4O)OC(C)C(C4O)O)3)c(c2)cc(c(c2OC)O)O)O)CO | + | SMILES C(C(O)6)(OC(C(O)C(O)6)Oc(c1)cc(c(C3=O)c1OC(=C(OC(C5O)OC(C(C(O)5)O)COC(C4O)OC(C)C(C4O)O)3)c(c2)cc(c(c2OC)O)O)O)CO |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
56 61 0 0 0 0 0 0 0 0999 V2000
-0.7143 0.9256 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7143 0.1006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0002 -0.3118 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7146 0.1005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7144 0.9255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0000 1.3380 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.4291 -0.3118 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.1434 0.1005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.1433 0.9255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.4288 1.3381 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.4291 -0.9550 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.8576 1.3379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5857 0.9175 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3138 1.3379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3138 2.1786 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5858 2.5991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.8576 2.1786 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4285 1.3379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.8985 -0.3385 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0003 -1.1365 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.0553 -2.6093 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2062 -2.7590 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.9206 -3.2419 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2753 -4.0332 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.1245 -3.8836 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.4100 -3.4005 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.5465 -2.0999 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.0418 -0.8823 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.1926 -1.0319 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.9071 -1.5148 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2618 -2.3061 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.1110 -2.1565 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3965 -1.6735 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5525 -0.4933 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.2560 -1.7008 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.3240 -2.8092 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.7919 -2.7480 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.5375 -3.2782 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9587 -2.1419 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9354 -3.2364 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.7180 -4.5008 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.8930 2.5663 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.9672 0.9665 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2508 1.3499 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.7300 0.6625 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.9803 0.9540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1988 0.9454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7825 1.4877 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.5484 1.2127 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.9190 1.0770 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.5088 0.6534 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.2397 0.4379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.5858 3.3485 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.1527 4.5008 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.0709 1.7353 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.3240 2.0167 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
1 18 1 0 0 0 0
8 19 1 0 0 0 0
3 20 1 0 0 0 0
21 22 1 0 0 0 0
22 23 1 1 0 0 0
23 24 1 1 0 0 0
25 24 1 1 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
21 27 1 0 0 0 0
28 29 1 0 0 0 0
29 30 1 1 0 0 0
30 31 1 1 0 0 0
32 31 1 1 0 0 0
32 33 1 0 0 0 0
33 28 1 0 0 0 0
28 34 1 0 0 0 0
33 35 1 0 0 0 0
32 36 1 0 0 0 0
31 37 1 0 0 0 0
25 38 1 0 0 0 0
37 38 1 0 0 0 0
22 39 1 0 0 0 0
23 40 1 0 0 0 0
24 41 1 0 0 0 0
29 19 1 0 0 0 0
42 15 1 0 0 0 0
14 43 1 0 0 0 0
44 45 1 1 0 0 0
45 46 1 1 0 0 0
47 46 1 1 0 0 0
47 48 1 0 0 0 0
48 49 1 0 0 0 0
49 44 1 0 0 0 0
44 50 1 0 0 0 0
45 51 1 0 0 0 0
46 52 1 0 0 0 0
47 18 1 0 0 0 0
53 54 1 0 0 0 0
16 53 1 0 0 0 0
55 56 1 0 0 0 0
49 55 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 53 54
M SBL 1 1 59
M SMT 1 OCH3
M SBV 1 59 0.0000 -0.7494
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 55 56
M SBL 2 1 61
M SMT 2 ^ CH2OH
M SBV 2 61 0.5225 -0.5225
S SKP 5
ID FL5FAHGL0004
FORMULA C34H42O22
EXACTMASS 802.216773028
AVERAGEMASS 802.68408
SMILES C(C(O)6)(OC(C(O)C(O)6)Oc(c1)cc(c(C3=O)c1OC(=C(OC(C5O)OC(C(C(O)5)O)COC(C4O)OC(C)C(C4O)O)3)c(c2)cc(c(c2OC)O)O)O)CO
M END