Mol:FL5FECGS0035
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 38 41 0 0 0 0 0 0 0 0999 V2000 | + | 38 41 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.4561 -3.7587 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.4561 -3.7587 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0361 -4.1716 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0361 -4.1716 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6397 -3.9518 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6397 -3.9518 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7513 -3.3193 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7513 -3.3193 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.2592 -2.9063 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.2592 -2.9063 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3445 -3.1260 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3445 -3.1260 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3548 -3.0995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3548 -3.0995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4665 -2.4670 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4665 -2.4670 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.9743 -2.0540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.9743 -2.0540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3707 -2.2737 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3707 -2.2737 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.7386 -3.4215 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.7386 -3.4215 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0858 -1.4216 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0858 -1.4216 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.7010 -1.1978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.7010 -1.1978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8147 -0.5530 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8147 -0.5530 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3131 -0.1322 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3131 -0.1322 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6980 -0.3563 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6980 -0.3563 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5843 -1.0008 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5843 -1.0008 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1316 -4.3646 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1316 -4.3646 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3848 1.5399 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 2.3848 1.5399 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.0552 0.8704 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.0552 0.8704 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.6625 1.1842 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6625 1.1842 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2445 1.0189 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.2445 1.0189 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.6640 1.5923 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.6640 1.5923 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.9668 1.3746 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 2.9668 1.3746 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 1.8261 1.3902 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8261 1.3902 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2050 1.7872 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2050 1.7872 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.4032 1.3942 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.4032 1.3942 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9559 0.6339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9559 0.6339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4388 0.2311 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4388 0.2311 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0352 1.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0352 1.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0750 -4.9661 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0750 -4.9661 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7002 -5.2482 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7002 -5.2482 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4062 -2.1250 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4062 -2.1250 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3459 -1.7830 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3459 -1.7830 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1597 -3.6346 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1597 -3.6346 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1445 -3.4610 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1445 -3.4610 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5489 0.8384 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5489 0.8384 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8976 0.0906 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8976 0.0906 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 19 20 1 0 0 0 0 | + | 19 20 1 0 0 0 0 |
| − | 20 21 1 1 0 0 0 | + | 20 21 1 1 0 0 0 |
| − | 21 22 1 1 0 0 0 | + | 21 22 1 1 0 0 0 |
| − | 23 22 1 1 0 0 0 | + | 23 22 1 1 0 0 0 |
| − | 23 24 1 0 0 0 0 | + | 23 24 1 0 0 0 0 |
| − | 24 19 1 0 0 0 0 | + | 24 19 1 0 0 0 0 |
| − | 19 25 1 0 0 0 0 | + | 19 25 1 0 0 0 0 |
| − | 24 26 1 0 0 0 0 | + | 24 26 1 0 0 0 0 |
| − | 23 27 1 0 0 0 0 | + | 23 27 1 0 0 0 0 |
| − | 20 28 1 0 0 0 0 | + | 20 28 1 0 0 0 0 |
| − | 15 28 1 0 0 0 0 | + | 15 28 1 0 0 0 0 |
| − | 16 29 1 0 0 0 0 | + | 16 29 1 0 0 0 0 |
| − | 29 30 1 0 0 0 0 | + | 29 30 1 0 0 0 0 |
| − | 2 31 1 0 0 0 0 | + | 2 31 1 0 0 0 0 |
| − | 31 32 1 0 0 0 0 | + | 31 32 1 0 0 0 0 |
| − | 8 33 1 0 0 0 0 | + | 8 33 1 0 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 1 35 1 0 0 0 0 | + | 1 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 22 37 1 0 0 0 0 | + | 22 37 1 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | M STY 1 5 SUP | + | M STY 1 5 SUP |
| − | M SLB 1 5 5 | + | M SLB 1 5 5 |
| − | M SAL 5 2 37 38 | + | M SAL 5 2 37 38 |
| − | M SBL 5 1 40 | + | M SBL 5 1 40 |
| − | M SMT 5 CH2OH | + | M SMT 5 CH2OH |
| − | M SVB 5 40 3.5974 1.0456 | + | M SVB 5 40 3.5974 1.0456 |
| − | M STY 1 4 SUP | + | M STY 1 4 SUP |
| − | M SLB 1 4 4 | + | M SLB 1 4 4 |
| − | M SAL 4 2 35 36 | + | M SAL 4 2 35 36 |
| − | M SBL 4 1 38 | + | M SBL 4 1 38 |
| − | M SMT 4 OCH3 | + | M SMT 4 OCH3 |
| − | M SVB 4 38 -3.4624 0.4373 | + | M SVB 4 38 -3.4624 0.4373 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 33 34 | + | M SAL 3 2 33 34 |
| − | M SBL 3 1 36 | + | M SBL 3 1 36 |
| − | M SMT 3 OCH3 | + | M SMT 3 OCH3 |
| − | M SVB 3 36 -0.6811 -1.0301 | + | M SVB 3 36 -0.6811 -1.0301 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 31 32 | + | M SAL 2 2 31 32 |
| − | M SBL 2 1 34 | + | M SBL 2 1 34 |
| − | M SMT 2 OCH3 | + | M SMT 2 OCH3 |
| − | M SVB 2 34 -3.6888 -1.3644 | + | M SVB 2 34 -3.6888 -1.3644 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 29 30 | + | M SAL 1 2 29 30 |
| − | M SBL 1 1 32 | + | M SBL 1 1 32 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SVB 1 32 0.5265 1.6977 | + | M SVB 1 32 0.5265 1.6977 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FECGS0035 | + | ID FL5FECGS0035 |
| − | KNApSAcK_ID C00005682 | + | KNApSAcK_ID C00005682 |
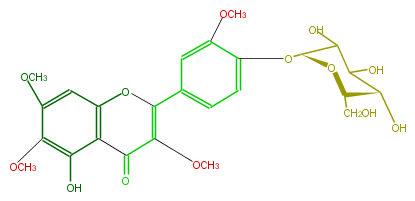
| − | NAME Chrysosplenin | + | NAME Chrysosplenin |
| − | CAS_RN 23279-19-8 | + | CAS_RN 23279-19-8 |
| − | FORMULA C25H28O13 | + | FORMULA C25H28O13 |
| − | EXACTMASS 536.152990982 | + | EXACTMASS 536.152990982 |
| − | AVERAGEMASS 536.48202 | + | AVERAGEMASS 536.48202 |
| − | SMILES C(C(OC)=2)(=O)c(c(OC2c(c3)ccc(O[C@@H](C4O)O[C@@H]([C@@H](C(O)4)O)CO)c(OC)3)1)c(O)c(OC)c(c1)OC | + | SMILES C(C(OC)=2)(=O)c(c(OC2c(c3)ccc(O[C@@H](C4O)O[C@@H]([C@@H](C(O)4)O)CO)c(OC)3)1)c(O)c(OC)c(c1)OC |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
38 41 0 0 0 0 0 0 0 0999 V2000
-0.4561 -3.7587 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0361 -4.1716 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6397 -3.9518 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7513 -3.3193 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2592 -2.9063 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3445 -3.1260 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3548 -3.0995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.4665 -2.4670 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.9743 -2.0540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3707 -2.2737 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.7386 -3.4215 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.0858 -1.4216 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.7010 -1.1978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8147 -0.5530 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3131 -0.1322 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6980 -0.3563 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5843 -1.0008 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1316 -4.3646 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.3848 1.5399 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.0552 0.8704 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.6625 1.1842 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.2445 1.0189 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.6640 1.5923 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.9668 1.3746 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
1.8261 1.3902 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.2050 1.7872 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.4032 1.3942 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9559 0.6339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.4388 0.2311 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0352 1.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0750 -4.9661 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.7002 -5.2482 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4062 -2.1250 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.3459 -1.7830 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1597 -3.6346 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.1445 -3.4610 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5489 0.8384 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.8976 0.0906 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
19 20 1 0 0 0 0
20 21 1 1 0 0 0
21 22 1 1 0 0 0
23 22 1 1 0 0 0
23 24 1 0 0 0 0
24 19 1 0 0 0 0
19 25 1 0 0 0 0
24 26 1 0 0 0 0
23 27 1 0 0 0 0
20 28 1 0 0 0 0
15 28 1 0 0 0 0
16 29 1 0 0 0 0
29 30 1 0 0 0 0
2 31 1 0 0 0 0
31 32 1 0 0 0 0
8 33 1 0 0 0 0
33 34 1 0 0 0 0
1 35 1 0 0 0 0
35 36 1 0 0 0 0
22 37 1 0 0 0 0
37 38 1 0 0 0 0
M STY 1 5 SUP
M SLB 1 5 5
M SAL 5 2 37 38
M SBL 5 1 40
M SMT 5 CH2OH
M SVB 5 40 3.5974 1.0456
M STY 1 4 SUP
M SLB 1 4 4
M SAL 4 2 35 36
M SBL 4 1 38
M SMT 4 OCH3
M SVB 4 38 -3.4624 0.4373
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 33 34
M SBL 3 1 36
M SMT 3 OCH3
M SVB 3 36 -0.6811 -1.0301
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 31 32
M SBL 2 1 34
M SMT 2 OCH3
M SVB 2 34 -3.6888 -1.3644
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 29 30
M SBL 1 1 32
M SMT 1 OCH3
M SVB 1 32 0.5265 1.6977
S SKP 8
ID FL5FECGS0035
KNApSAcK_ID C00005682
NAME Chrysosplenin
CAS_RN 23279-19-8
FORMULA C25H28O13
EXACTMASS 536.152990982
AVERAGEMASS 536.48202
SMILES C(C(OC)=2)(=O)c(c(OC2c(c3)ccc(O[C@@H](C4O)O[C@@H]([C@@H](C(O)4)O)CO)c(OC)3)1)c(O)c(OC)c(c1)OC
M END